Compare actin filament compression between different fiber-scale simulators
We have previously compared simulations of single actin filament compression at monomer- and fiber-scales using the ReaDDy and Cytosim simulation engines, respectively, under different compression velocities. To better understand how these difference we observed may be specific to the choice of simulators, we proposed running an additional comparison between two simulators at fiber scale.
Interactive visualizations of the previous simulations are available at here.
Proposed by:
Megan Riel-Mehan
Publication
https://www.micropublication.org/journals/biology/micropub-biology-001347
Preliminary Findings
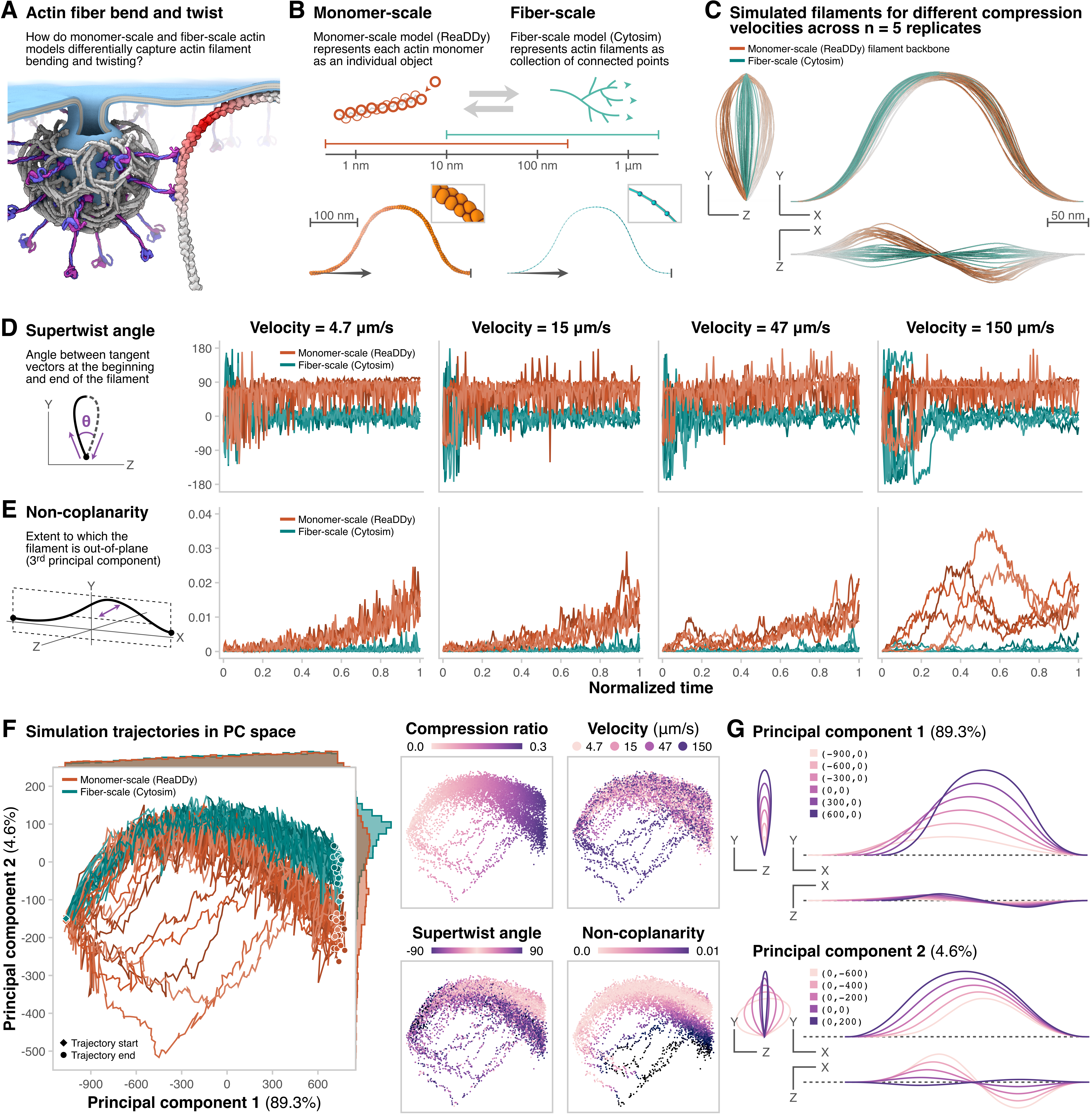
Fiber-scale simulations of single actin filament compression (using Cytosim) were not able to capture filament supertwist behavior observed with monomer-scale simulations (using ReaDDy).

Suggested next steps:
- Build a MEDYAN model for compression of a single actin filament
- Run simulations at different compression velocities
- Use the subcell-pipeline to align filaments, perform dimensionality reduction, and calculate metrics
Resources
a description for the tool
Materials and methods available:
- actin
- computational modeling